An integration layer for neural simulation:
PyNN in the software forest



















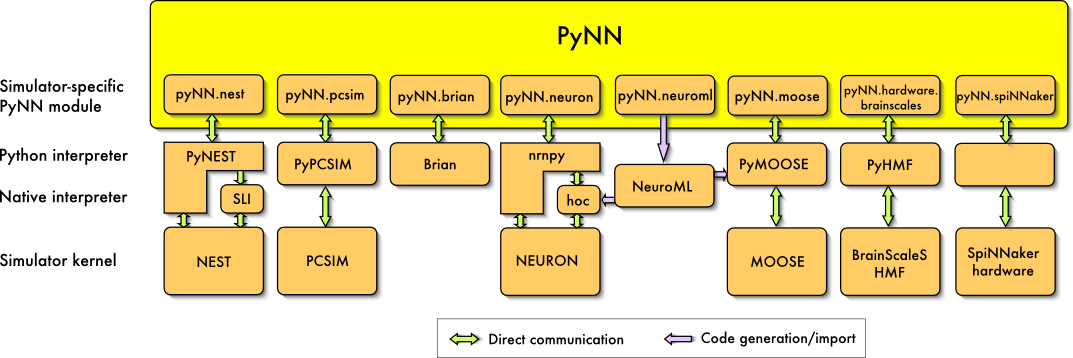
PyNN (pronounced 'pine') is a simulator-independent language for building spiking neuronal network models. A PyNN script runs without modification on any simulator or neuromorphic hardware platform that PyNN supports (currently NEURON, NEST, PCSIM, MOOSE, Brian, the BrainScaleS platform and the SpiNNaker platform).
PyNN provides a library of standard neuron, synapse and synaptic plasticity models, which have been verified to work the same on the different supported simulators. PyNN also provides a set of commonly-used connectivity algorithms (e.g. all-to-all, random, distance-dependent, small-world) but makes it easy to provide your own connectivity in a simulator-independent way.
Even if you don't wish to run simulations on multiple simulators, you may benefit from writing your simulation code using PyNN's powerful, high-level interface. In this case, you can use any neuron or synapse model supported by your simulator, and are not restricted to the standard models.
PyNN is open-source software (CeCILL licence). For downloads and full documentation see http://neuralensemble.org/PyNN/. For more information, see Davison et al. (2009)
In this poster, we showcase five recent developments that extend PyNN's capabilities. In each case, the extension involves integrating PyNN with existing, stand-alone software. Taken as a whole, these new features are a good illustration both of the merits of Python in general and PyNN in particular as a federation platform/integration tool for neuronal simulation (cf Cornelis et al. 2012), and of the benefits of a modular approach to neuroscience software development.

NeuroML (Gleeson et al., 2010) is an XML-based, declarative language for describing neuronal network models, in particular models with detailed dendritic morphologies. Exporting PyNN models to NeuroML is very straightforward:
import pyNN.nineml as sim
sim.setup(file="my_model.nml")
# the rest of your script as usual.
Only the model description is exported. The description of what to record, how long to simulate for, etc. is not exported. We are looking into using the SED-ML language to export this information. Importing NeuroML descriptions into PyNN is planned for the near future.

NineML (Raikov et al., 2011) is another XML-based, declarative language for describing neuronal network models. It was developed to support explicit specification of the mathematics of neuronal and synaptic models, in contrast to the implicit specification in NeuroML v1 (note however, that NeuroML v2 will also support such explicit mathematical specifications). For example, here is an integrate-and-fire neuron described using the Python interface to NineML:
import nineml.abstraction_layer as al
iaf = al.ComponentClass(name="iaf_tau",
regimes=[al.Regime(name="subthreshold_regime",
time_derivatives=["dv/dt = (v_rest - v)/tau_m + (i_offset + i_syn)/cm",],
transitions=al.On("v > v_thresh",
do=["t_spike = t", "v = v_reset",
al.OutputEvent('spike_output')],
to="refractory_regime"),),
al.Regime(name="refractory_regime",
time_derivatives=["dv/dt = 0"],
transitions=[al.On("t >= t_spike + tau_refrac", to="subthreshold_regime")])],
state_variables=[al.StateVariable('v'), al.StateVariable('t_spike')],
analog_ports=[al.SendPort("v"), al.ReducePort("i_syn", reduce_op="+")],
event_ports=[al.SendEventPort('spike_output')])
Where the backend supports it (currently NEURON, with NEST & Brian support under development), you can directly use such NineML descriptions in PyNN:
import pyNN.neuron as sim
celltype_cls = sim.nineml.nineml_celltype_from_model(name="iaf", nineml_model=iaf)
cells = sim.Population(1000, celltype_cls)
As for NeuroML, we are currently working on exporting and importing entire PyNN scripts as NineML.

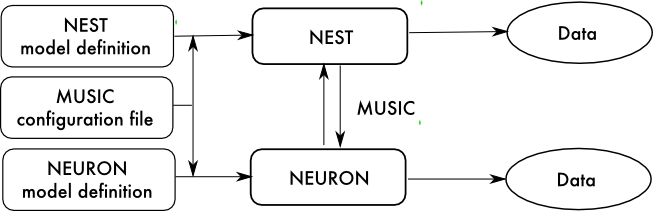

The MUSIC library Djurfeldt et al., 2010) enables independent, parallel neural simulators to exchange data, such as spike times, and as such greatly facilitates multi-scale modelling.
There remain several barriers to routine use of such multi-simulations. In particular, each part of the model must be programmed using the particular language used by the simulator on which it will be simulated, which can make coordination difficult. Similarly, each simulator will export data in its own particular format, and manipulation of such heterogeneous datasets to enable a coherent analysis can be equally challenging.
By combining PyNN and MUSIC, we gain the ability to specify a multi-simulator model and experimental protocol in a single script using a single programming language. In other words, MUSIC adds to PyNN the ability to distribute a model over multiple simulators, while PyNN adds to MUSIC the possibility of describing a multi-simulator model in a single script, and a unified representation of output data.
from pyNN import music
sim1, sim2 = music.setup(music.Config("nest", 1), music.Config("neuron", 1))
sim1.setup(timestep=0.1, min_delay=0.5)
sim2.setup(timestep=0.05, min_delay=0.5)
output_cell_type = sim2.IF_curr_alpha(tau_refrac=2.0, v_thresh=-50.0, tau_syn_E=2.0, tau_syn_I=2.0)
output_population = sim2.Population(2, output_cell_type, label="output")
input_population = sim1.Population(1, sim1.SpikeSourcePoisson(rate=100.0), label="input")
projection = music.Projection(input_population, output_population, sim2.AllToAllConnector(),
sim2.StaticSynapse(weight=1.0, delay=0.5))
input_population.record('spikes')
output_population.record(['spikes', 'v'])
music.run(1000.0)
input_population.write_data("Results/simpleNetwork_input_%s.pkl" % sim1.name)
output_population.write_data("Results/simpleNetwork_output_%s.pkl" % sim2.name)
music.end()
Neo is a package for representing electrophysiology data in Python, together with support for reading a wide range of neurophysiology file formats, including Spike2, NeuroExplorer, AlphaOmega, Axon, Blackrock, Plexon, Tdt, and support for writing to a subset of these formats plus other formats including Matlab and HDF5.
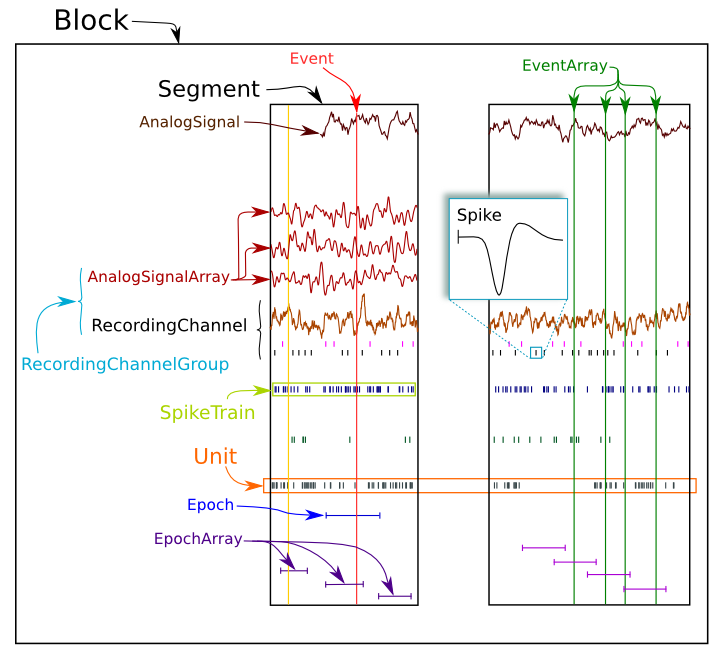
The goal of Neo is to improve interoperability between Python tools for analyzing, visualizing and generating electrophysiology data by providing a common, shared object model. Neo implements a hierarchical data model well adapted to intracellular and extracellular electrophysiology and EEG data with support for multi-electrodes. Neo objects behave just like normal NumPy arrays, but with additional metadata, checks for dimensional consistency and automatic unit conversion.
Using Neo provides several advantages:
reset() between each one);Here is a very simple example:
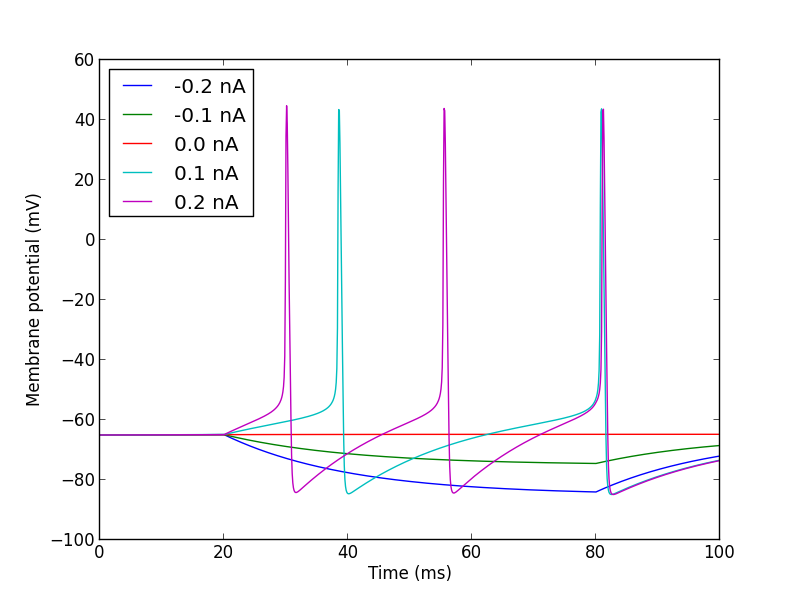
import pyNN.neuron as sim
import matplotlib.pyplot as plt
from quantities import nA
sim.setup()
cell = sim.Population(1, sim.HH_cond_exp())
step_current = sim.DCSource(start=20.0, stop=80.0)
step_current.inject_into(cell)
cell.record('v')
for amp in (-0.2, -0.1, 0.0, 0.1, 0.2):
step_current.amplitude = amp
sim.run(100.0)
sim.reset(annotations={"amplitude": amp*nA})
data = cell.get_data()
sim.end()
for segment in data.segments:
vm = segment.analogsignalarrays[0]
plt.plot(vm.times, vm,
label=str(segment.annotations["amplitude"]))
plt.legend(loc="upper left")
plt.xlabel("Time (%s)" % vm.times.units._dimensionality)
plt.ylabel("Membrane potential (%s)" % vm.units._dimensionality)
plt.show()

The connection-set algebra (CSA; Djurfeldt, 2012) is a general formalism for specifying the connectivity of neuronal network models. CSA provides operators to form more complex sets of connections from simpler ones. The CSA is expressive enough to describe a wide range of connectivities and can serve as a concise notation for network structure in scientific writing as well as in model description code for neural simulations.
For example, random connectivity is represented by \(\overline{\rho}\), while one-to-one connections are represented by \(\overline{\delta}\). To connect a population randomly to itself with 50% connectivity, but without autapses, you would use \(\overline{\rho}(0.5) - \overline{\delta}\). Much more complex connectivity, including dependencies on spatial geometry, can also easily be expressed.
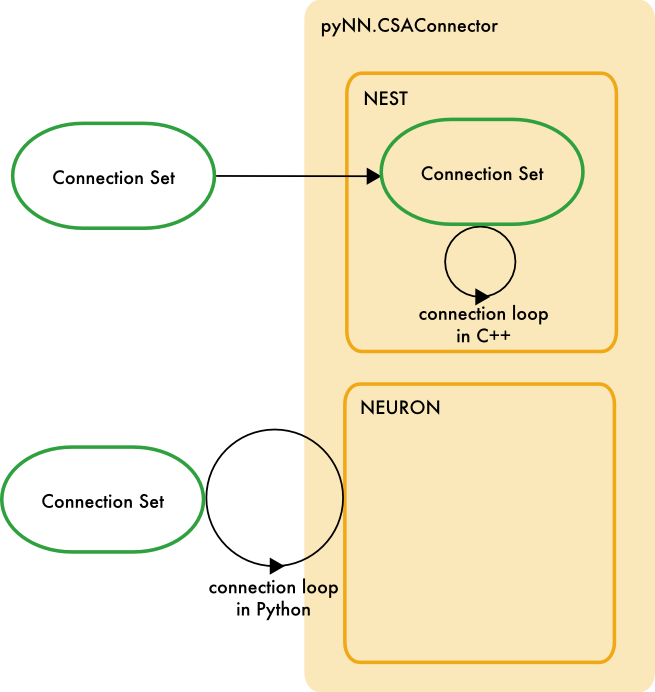
PyNN provides the CSAConnector class, which accepts a ConnectionSet object from the Python csa library, and connects a PyNN network accordingly:
import csa
import pyNN.nest as sim
from csa import cset, random, oneToOne
...
connection_set = cset(random(0.5) - oneToOne, weight, delay)
projection = sim.Projection(pre, post,
sim.CSAConnector(connection_set))
...
Where the backend simulator supports the ConnectionGenerator interface, PyNN passes the ConnectionSet object to the backend, where the connection loop can take place in C++, for much improved performance. Otherwise, the connection loop takes place within PyNN.